Protein Blocks¶
Protein Blocks (PBs) are structural prototypes defined by de Brevern et al [1]. The 3-dimensional local structure of a protein backbone can be modelized as an 1-dimensional sequence of PBs. In principle, any conformation of any amino acid could be represented by one of the sixteen available Protein Blocks.
PBs are labeled from a to p (see Figure 1). PBs m and d can be roughly described as prototypes for alpha-helix and central beta-strand, respectively. PBs a to c primarily represent beta-strand N-caps and PBs e and f, beta-strand C-caps; PBs a to j are specific to coils, PBs k and l to alpha-helix N-caps, and PBs n to p to alpha-helix C-caps.
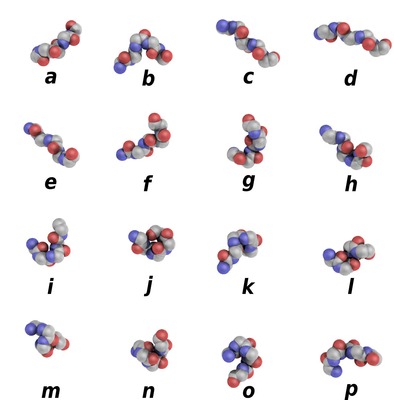
Figure 1. Schematic representation of the sixteen protein blocks, labeled from a to p (Creative commons CC-BY).
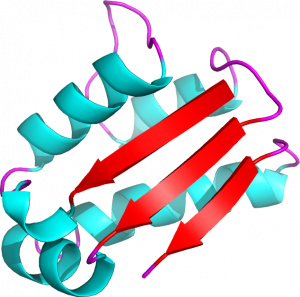
Figure 2. 3D representation of the barstar protein (PDB ID 1AY7, chain B) (Creative commons CC-BY).
For instance, the 3D-structure of the barstar protein represented in Figure 2 can be translated into a 1D-sequence of PBs:
ZZdddfklpcbfklmmmmmmmmnopafklgoiaklmmmmmmmmpacddddddehkllmmmmnnommmmmmmmmmmmmmnopacddddZZ
The conformations of the 89 residues of the barstar are translated into a sequence of 89 PBs. Note that “Z” corresponds to amino acids for which a PB cannot be assigned. As a matter of fact, the assignment of a given residue n requires is based on the conformations of residues n-2, n-1, n, n+1 and n+2. Therefore, no PB can be assigned to the two first (N-termini) and two last (C-termini) residues of a polypeptide chain.
Neq¶
Neq is calculated as follows:
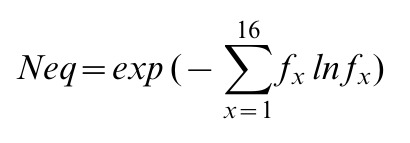
Where fx is the probability of PB x. Neq quantifies the average number of PBs at a given position in the protein sequence. A Neq value of 1 indicates that only one type of PB is observed. On the opposite, a Neq value of 16 is equivalent to a fully random distribution of PBs.
| [1] |
|